Pharmacogenomics: A personalized response to medication non-adherence
- Pharmacogenetics or the genetic variations in the human population giving rise to a variable drug response is well-known in clinical research1.
- The quest for personalized medicine based on pharmacogenomics using an individual’s genetic content has gained traction since the completion of the first draft of the human genome project2,3.
- Pharmacogenomics-based personalized prescriptions can help in improving treatment efficacy and reducing drug-related side effects, the two major barriers to medication adherence1.
- However major hurdle remains in the widespread implementation of pharmacogenetic testing that needs to be addressed before it can be employed to improve adherence in routine clinical practice1,2.
Medication non-adherence is a multifacet issue with enormous clinical and economical implications requiring a similarly multipronged solution. Several behavioral, educational, combinatorial, and smart interventions are employed to encourage adherence to the prescribed therapy (See: Interventions to tackle medication non-adherence; Smart solutions to the complicated problem of non-adherence). However, if patients don’t set to benefit from the treatment, they tend to become non-adherent even upon employing the best of these interventions. One of the reasons for this sub-optimal efficacy of a drug is individual differences in response to treatment owing to differential genetic makeup. This further enhances adverse effects, costs, and wastage of medical resources. The field of pharmacogenomics is trying to address this by bringing a personalized approach toward medication prescription. Prescriptions based on pharmacogenomics information can encourage medication adherence in patients because if they know the drug is tailor-made for them, they tend to adhere better1,2.
What are Pharmacogenetics and pharmacogenomics?
The study of genetic variations in the human population responsible for a variable drug response is called pharmacogenetics2. Decades of research have given a sneak peek into why certain individuals are resistant to life-threatening infections or why certain drugs work only in certain patients and not for others.
Completion of the first draft of the human genome project at the turn of this century was the first step toward personalized medicine. Although 99.9% of the human genome is identical, individuals differ from each other in 0.1% (1 in 1000 bases) of their genetic content. Most of these differences comprise variations in single base pairs, also called single nucleotide polymorphisms or SNPs4 that are responsible for our individualized response to a given treatment2. In the post-genomic era, pharmacogenetics has evolved more into pharmacogenomics. It focuses on a genome-wide study of factors responsible for variable drug metabolism, drug targets, or drug transport2. Since many chemotherapeutic targets are heritable traits, the existing genetic variants contribute to variable drug response1-3. The resulting information could be used for a personalized approach toward medicine3.
Pharmacogenetic testing in therapy management
Pharmacogenetic testing is at the forefront of precision medicine. It is estimated that 20-95% variability in individual response to drugs can be attributed to the genetic components1. Knowledge about a patient’s genetic predisposition that dictates treatment efficacy or the likelihood of drug-associated adverse effects is the cornerstone for providing a tailor-made therapy. For example, it is advised to test for HLA-B*57:01 before prescribing abacavir – an antiretroviral for HIV management – to identify patients at risk of developing severe adverse effects5. Several such gene-drug interacting pairs adapted for personalized therapy exist, a few are listed below:
- Phenytoin: An anticonvulsant drug used to treat seizures. Patients carrying a variation in human leucocyte antigen B and hepatic CYP2C9 enzyme are more prone to develop adverse effects. This supports prescribing an alternative to phenytoin in HLA-B variants and reducing the dosage in CYP2C9 variants for improving adherence6
- Sulphonylureas: Used for diabetes type 2 treatment via aiding insulin secretion by β-cells. Patients carrying variations in transcription factor TCF7L2 and sulphonylurea receptor ABC8 show poor response to sulphonylureas supporting alternative drug prescriptions for diabetes management7,8
- Statins: Statins reduce the risk of cardiovascular diseases. Genetic variants of Kinesin motor KIF6 display better response to intensive statin therapy9 and pharmacogenetic testing for KIF6 is reported to improve adherence10
- Warfarin: An anticoagulant prescribed for cardiovascular diseases. Variants for CYP2C9 are highly sensitive to warfarin doses resulting in excessive bleeding. While variants of VKORC1 show poor response to warfarin treatment11.
- Clopidrogel: Another anticoagulant prescribed for cardiovascular diseases shows reduced response in CYP2C19 variants12.
- Codeine: Prescribed for severe cough relief in patients. Patients carrying CYP2D6 gene variants can rapidly metabolize it into morphine causing drug toxicity and even death5.
Barriers to implementation of pharmacogenetic testing:
Even with vast scientific evidence about the benefits of genetic testing for personalized therapy, it is not widely used. Several barriers exist in the implementation of pharmacogenetic testing2 such as:
Physician awareness: Clinician awareness about pharmacogenomics testing for various drug treatments are inadequate. This includes limited knowledge about test availability, result interpretation, and context-specific application in therapy management2,13.
Poor test availability: Genetic testing involves specialized instruments and trained professionals to oversee instrumentation, sample preparation, result interpretation, and report preparation. Therefore genetic testing remains poorly available in routine clinical practice.
Higher Costs: Costs are the major impediment in pharmacogenetic testing implementation. Although the costs have been reduced significantly in the last decade, it still higher than regular biochemical tests. This could be countered by incentivizing genetic testing and including them in medical insurance claims.
Lack of guidelines: Limited guidelines exist for interpreting genetic testing results and integrating them into the decision-making process. Forums like the clinical pharmacogenetics implementation consortium (CPIC) are working to produce specific peer-reviewed guidelines for recommending personalized therapy based on pharmacogenetic testing13.
Limited clinical decision support: Clinicians need constant support in the decision-making process to utilize results from pharmacogenetic tests. This would need integrating electronic health records with updated data from different platforms documenting the diverse genotype-to-phenotype relationship13,14. A few databases that document the genetic variations associated with drug response and adverse effects are the pharmacogenomic knowledge base (PharmGKB), pharmacogenomics research network (PGRN), and pharmacogenetics implementation consortium (CPIC). However data linkage and updated documention of the full diversity of clinical response to all the new variants is still a major challenge15.
Patient’s negative perception: Certain patients refuse genetic testing due to their false perceptions about the tests, their utility, higher costs, and adverse effects. This issue is higher in patients with lower health literacy1.
Pharmacogenetics – a promise of personalized response to non-adherence
Slowly pharmacogenetics is moving from specialized therapies like cancers to more generally prescribed treatments like statins, warfarin, etc. At present, labels for about 113 FDA-approved drugs carry information for >500 pharmacogenetic biomarkers spanning different therapeutic areas1 listed in figure 1.
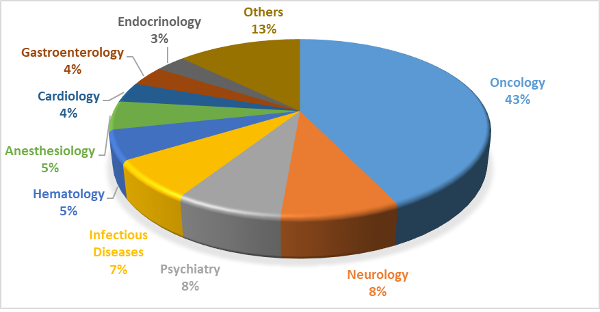
Although improving efficacy and reducing adverse effects is the primary goal of pharmacogenetic testing, they can play a major role in enhancing adherence1.
Information derived from pharmacogenetic testing could help in strengthening patients’ beliefs, a key factor in medication adherence5. It can help in reducing non-adherence in the following manner:
- Mitigate concerns for the therapy: Patients are aware of their odds of experiencing adverse effects, therefore are more confident about the treatment.
- Reinforce the need for therapy: Therapy is personalized for individuals and therefore they understand the importance of the treatment.
- Engage patients in decision-making: Patients also feel more involved in the decision-making process as the therapy is tailor-made for them.
- Build clinician-patient trust: As patients feel the clinicians are paying individualized attention, they trust their healthcare practitioners more.
- Reduce overall costs: Pharmacogenetic testing may be a bit higher initially, but overall it helps reduces healthcare costs by enhancing treatment effectiveness and reducing rehospitalizations due to adverse effects.
- Generate health literacy: Discussionsabout genetic factors and their influence on treatment effectiveness can enhance health literacy of the patients1.
- Improved outcomes in polypharmacy: A better understanding of the patient’s drug response, risk of adverse effects, and potential drug-drug interaction, pharmacogenetic testing can help in medication management in patients using multiple medications. This is especially the case in older age group suffering from multiple chronic conditions and find it difficult to adhere to their medications due to polypharmacy17.
Conclusion:
It is an established fact that medicines display variable efficacy and adverse effects in patients owing to variances in their response to a given drug which in turn is dictated by differences in their genetic makeup. Using pharamacogenomics to prescribe medicines can help in enhancing adherence to therapy essentially by influencing necessity vs. concern beliefs in patients. Combining pharmacogenetic information with environmental factors and patient’s clinical status will be required for a successful approach towards personalized medicine and for encouraging adherence1.
“In pharmacology it’s so clear that even if you have a fixed dose of a drug, the individuals respond very differently to one and the same dose”.
– By Arvid Carlsson, Neuroscientist and Nobel laureate













